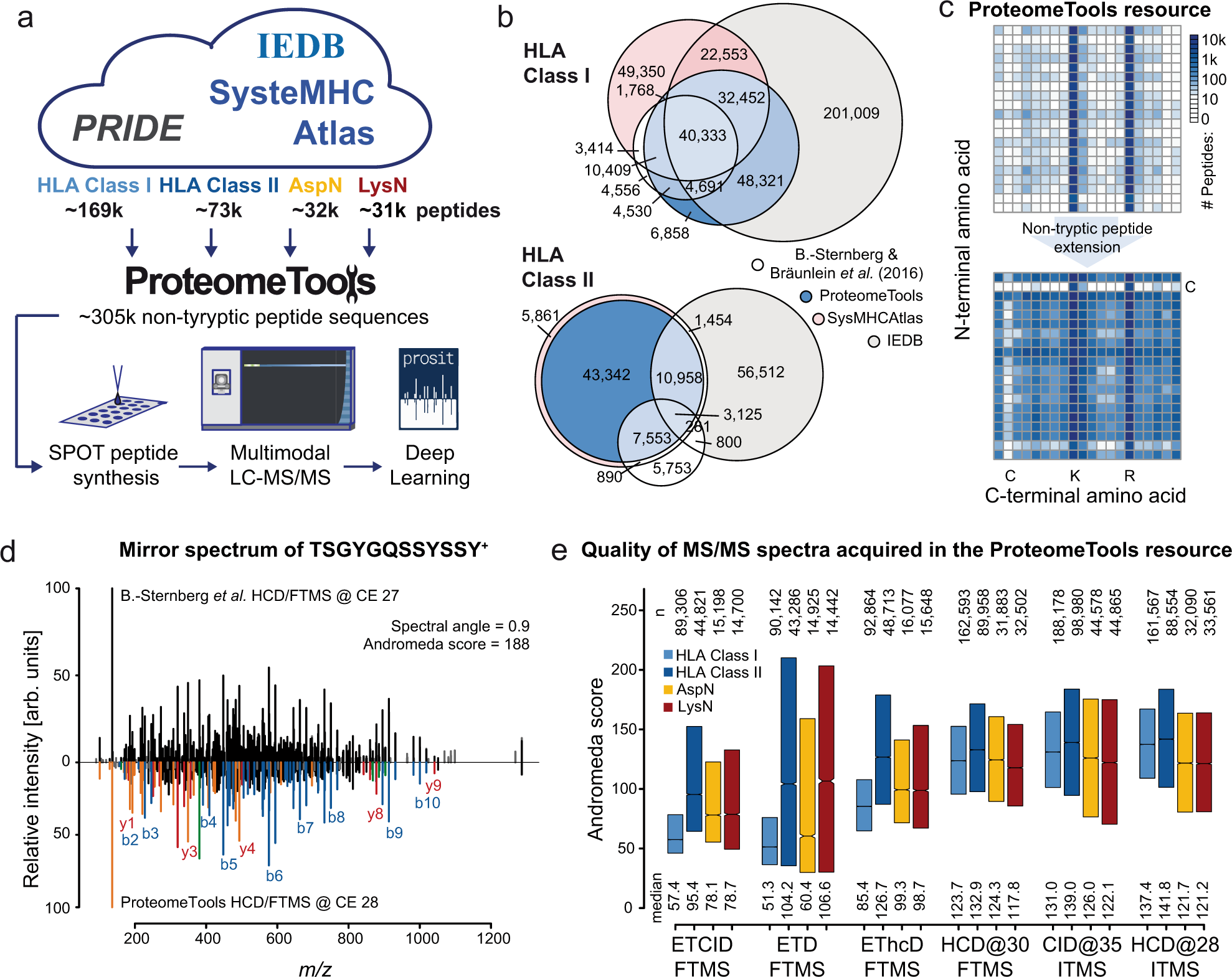
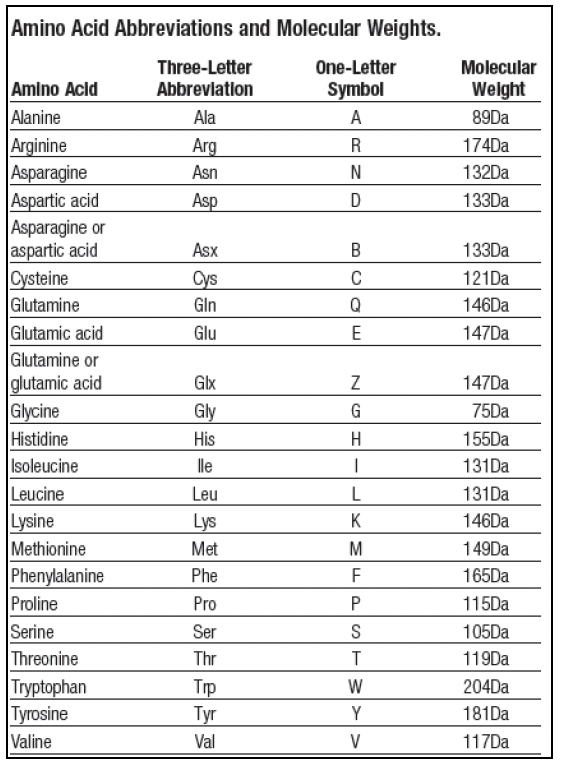
Functional expression of diverse post-translational peptide-modifying enzymes in Escherichia coli under uniform expression and purification conditions | PLOS ONE

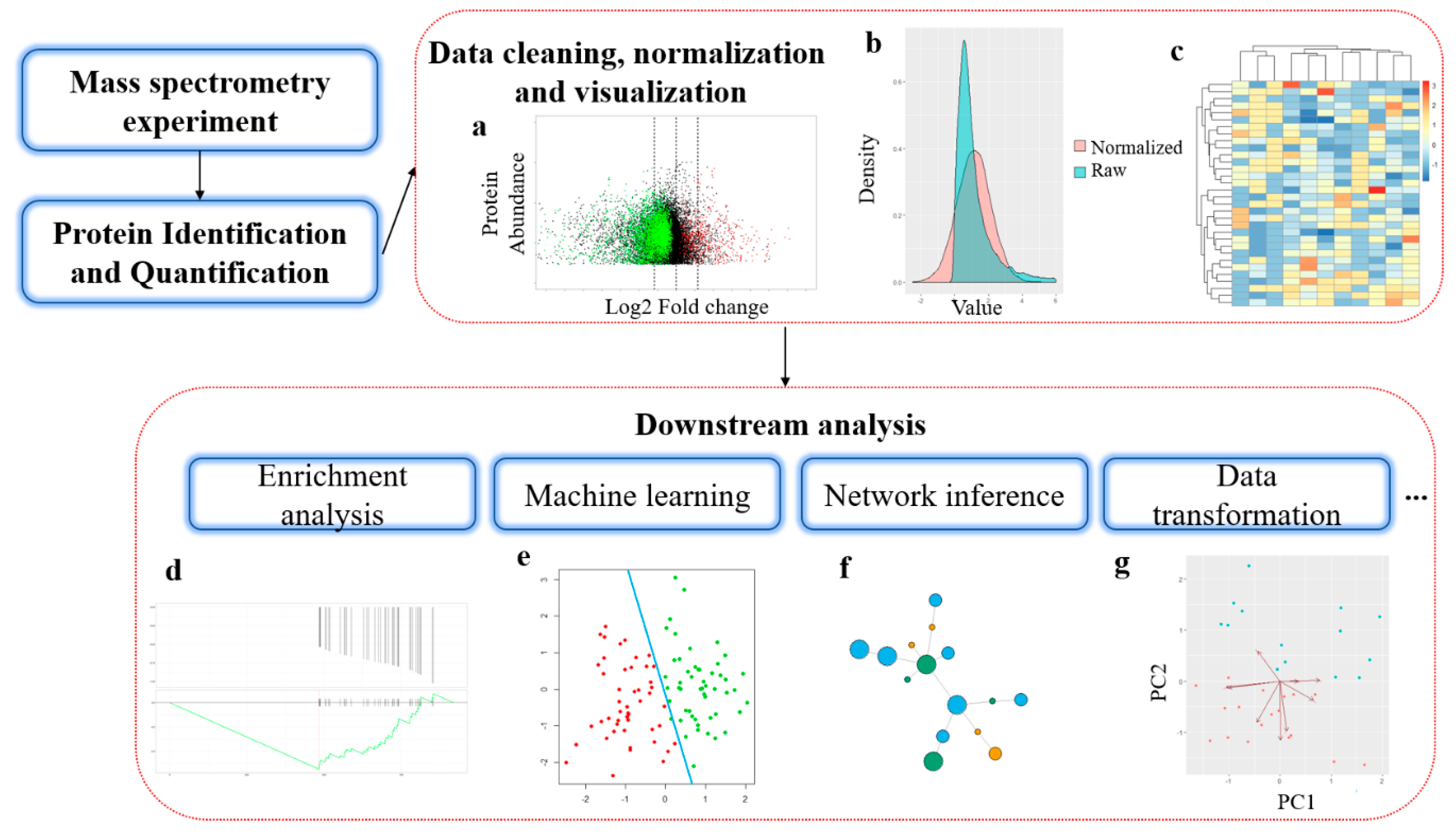
Data acquisition and database search. For a given experimental MS/MS... | Download Scientific Diagram

Computational generation of proteins with predetermined three-dimensional shapes using ProteinSolver - ScienceDirect

Protein structure, amino acid composition and sequence determine proteome vulnerability to oxidation‐induced damage | The EMBO Journal
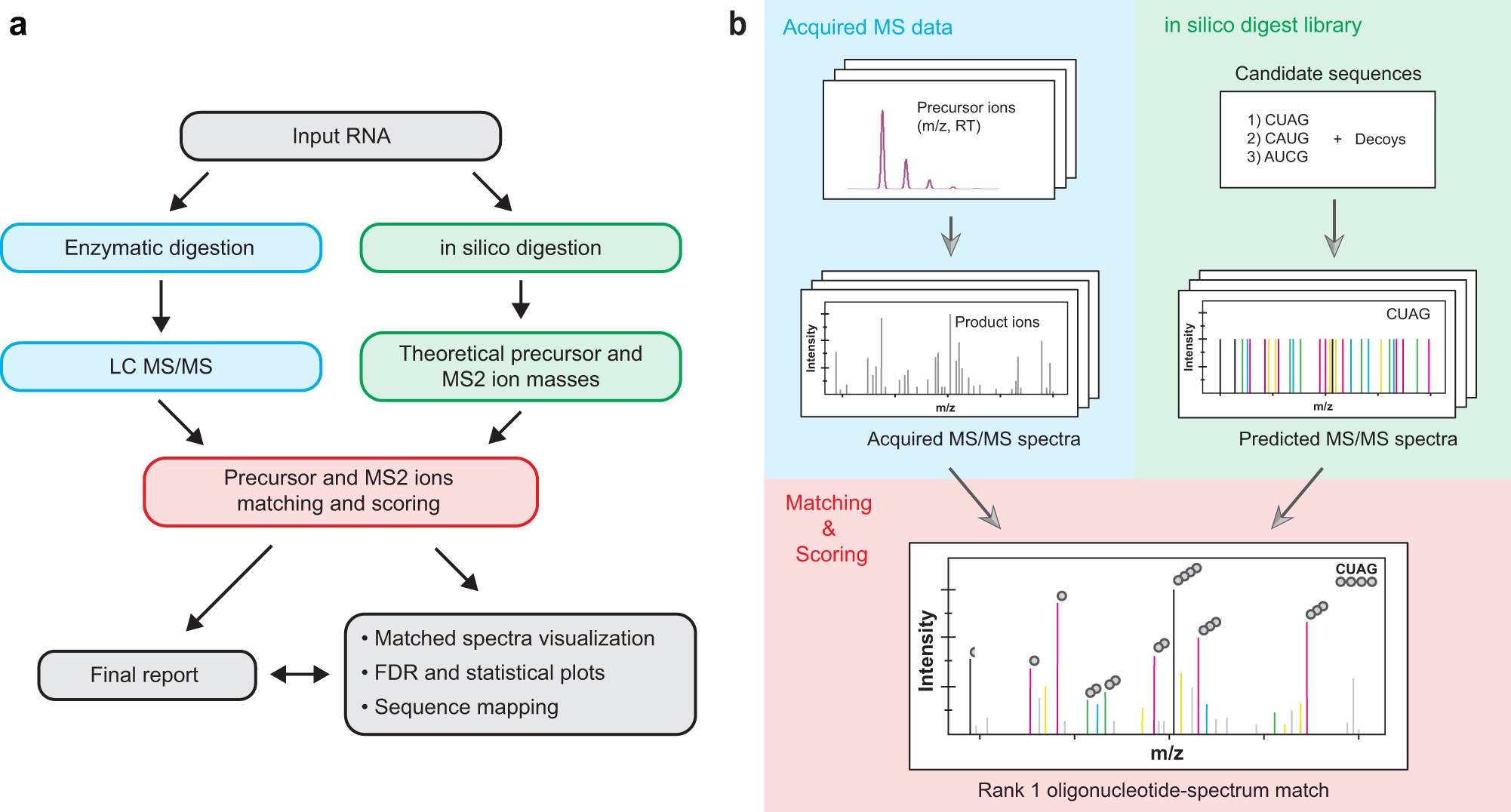
Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry | Nature Communications
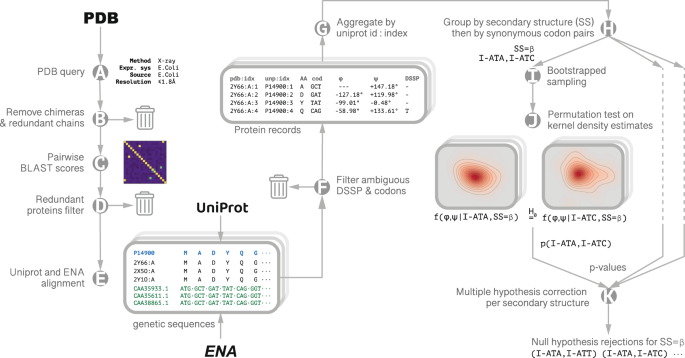
Codon-specific Ramachandran plots show amino acid backbone conformation depends on identity of the translated codon | Nature Communications

Frontiers | Essentially Leading Antibody Production: An Investigation of Amino Acids, Myeloma, and Natural V-Region Signal Peptides in Producing Pertuzumab and Trastuzumab Variants

Molecules | Free Full-Text | PepFun: Open Source Protocols for Peptide-Related Computational Analysis